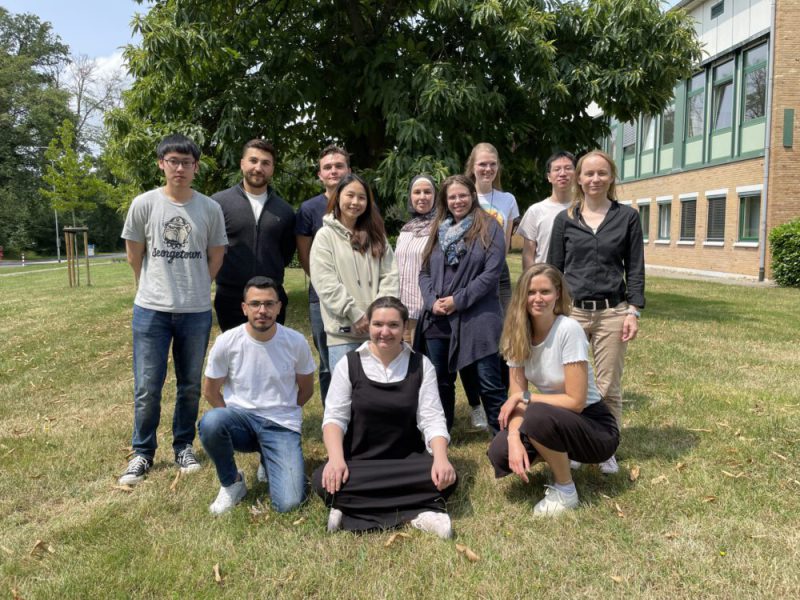
Discover the world of Molecular Modeling
Are you intrigued by the fascinating possibilities of molecular modeling techniques, such as molecular docking or molecular dynamics (MD) simulations? We, as a computational biochemistry research group, are happy to share our knowledge, either in our on-site BioSim course at the Heinrich-Heine University, through our comprehensive online MD Tutorial or by joining our group for an internship
KI in der Chemie und Biochemie (KIChem)
Inhalte des Moduls:
- Python in der Chemie
- Einführung in das Maschinelle Lernen (ML)
- Evaluierung und Interpretation von ML-Modellen
- Typische KI/ML-Programme in der (Bio-)Chemie
Vorlesung: Dienstags, 14:30–16:15 (ab 08.04.2025), Hörsaal 6F
Computerübungen: Mittwochs 14:30–16:00 oder 16:30–18:00 oder Freitags: 14:30–16:00
Abschlussprüfung: Klausur (8 ECTS-Punkte)
Ort: Heinrich Heine Universität, Raum: 25.41.00.43
Dozentin: Prof. Dr. Birgit Strodel & Tutor*innen
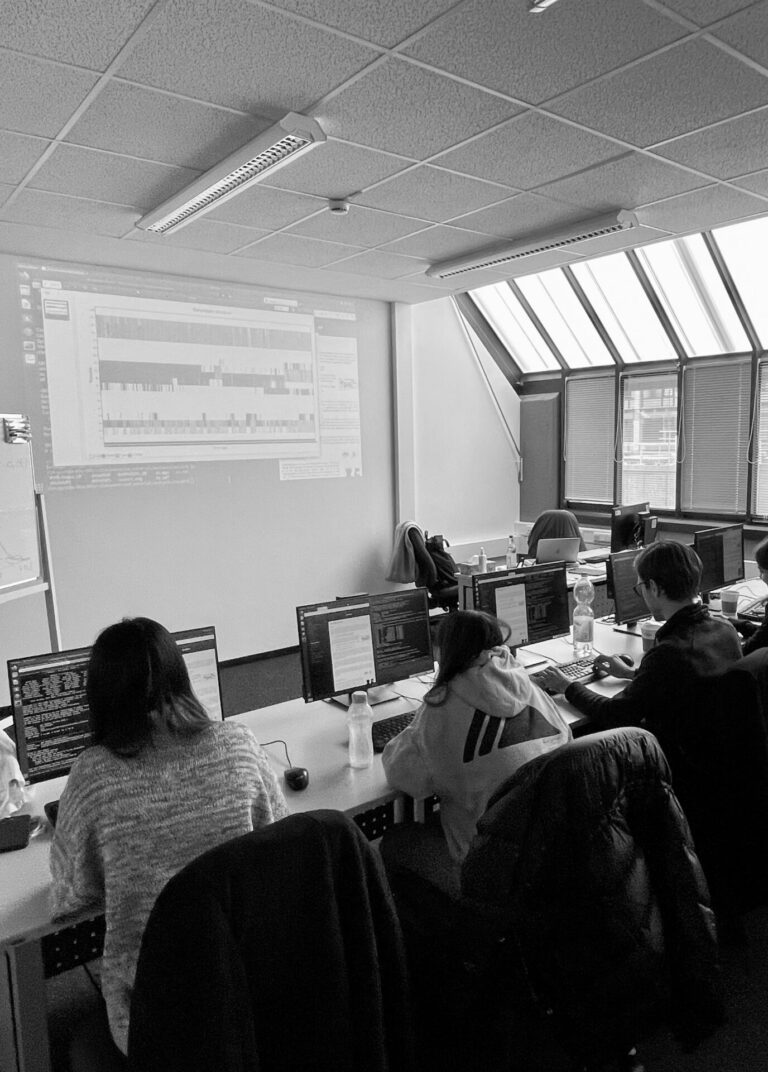

Simulation of Biomolecules (BioSim) course
The next BioSim course will take place in March 2026!
Our BioSim course aims to provide hands-on training in simulating biomolecules. You will learn how to set up and conduct MD simulations using the powerful and freely available MD software package, GROMACS.
When: Two and a half weeks in March, from 9 am to 5 pm with lectures in the morning and practical sessions in the afternoon
How: You can apply to this course through the HIS-LSF system at the Heinrich-Heine University or by contacting us directly
What to bring: None, computers are provided on-site
Online Tutorial for molecular dynamics (MD) simulations with GROMACS
For those eager to dive into MD simulations but seeking guidance, we’ve developed an extensive online MD tutorial using GROMACS. Our tutorial provides a structured introduction to the subject, walking you through every step of preparing and conducting an MD simulation.
In our MD tutorial, we present a practical example involving on the peptide [KIGAKI]3, which consists of 18 amino acids and serves as a model for studying protein aggregation.
⊕ Simulation setup
⊕ Energy minimization
⊕ Equilibration
⊕ MD Production run
⊕ A comprehensive selection of analysis methods
⊕ Enhanced sampling methods, such as replica exchange MD simulation
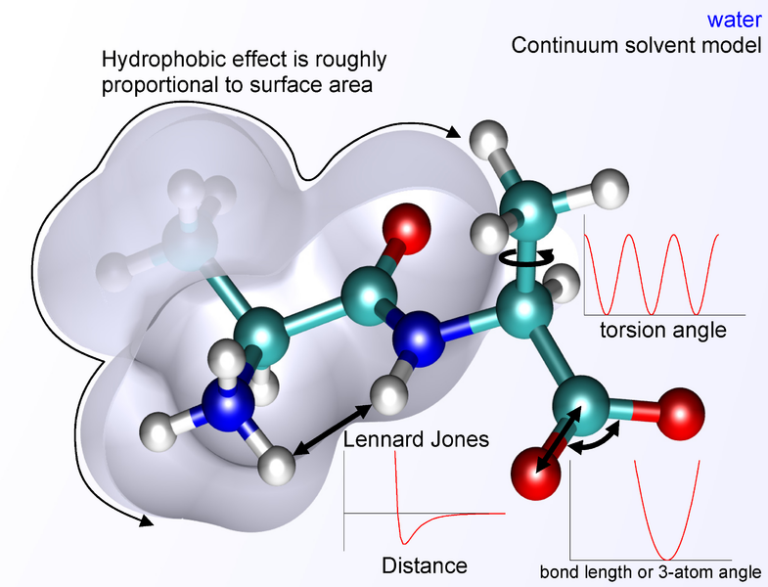
To get started, you’ll need a computer or laptop with a Linux (Ubuntu, openSUSE, etc.) or MacOS operating system, as GROMACS operates through a terminal interface. If you’re on Windows, you can use a virtual box with a Linux system.
Recommended Reading
If you want to learn more about the theory and practical applications of MD simulations, we recommend the following books:
- “Understanding Molecular Simulation” by Daan Frenkel and Berend Smit (1996)
- “Molecular Modelling: Principles and Applications” by Andrew R. Leach (2001)
- “Molecular Modeling and Simulation” by Tamar Schlick (2002)
Become part of our group
For those who cannot get enough of computational biochemistry, we also offer the opportunity to engage in an internship within our group, lasting from one to three months. During this immersive experience, you won’t just be learning about MD simulations – you will be actively involved in exciting research projects, applying your knowledge to practical scenarios, and gaining experience in this dynamic field.
This internship, or the successful completion of our BioSim course, can pave the way for those interested in writing their bachelor’s or master’s theses within our group. To learn more about our ongoing research, please visit our research site for an in-depth look into our projects and achievements.

